Dynamical systems play a key role across many application areas such as automotives, process industry, finance, biology, and aerospace. Such systems can be described by differential or difference equations, which describe the evolution of variables characterizing the system over time.
Details
We are developing tools and techniques to support the the model building process for dynamical systems. This involves modeling methodology using ordinary and stochastic differential equations, maximum likelihood based parameter estimation, Bayesian inference, and identifiability analysis.
Applications
Within a series of EU funded projects (YSBN, UNICELLSYS, SYSINBIO) we have applied dynamic systems modeling to biochemical reaction networks to model signaling and metabolic pathways. We are currently involved in IMPACT – an EU funded Marie Curie European Industrial Doctorate scheme where dynamic systems modeling of pharmacokinetic and pharmacodynamic processes is the common theme for five early stage researchers carrying out joint academic-industrial collaborative PhD-projects.
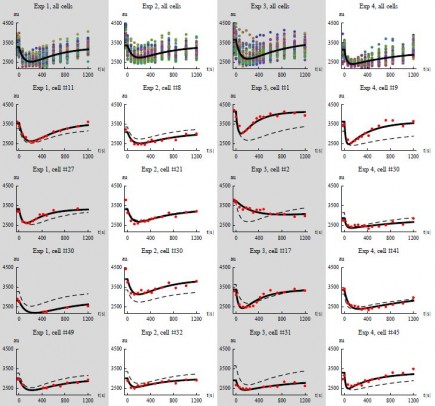
In a recently initiated project funded by SSF we address three application areas each of which can be described as dynamical systems, where tools and techniques for system identification and mixed effects modeling will be extended: proliferation and migration dynamics in brain tumor cells, human lipid metabolism in metabolic diseases, and regulation of glucose metabolism in yeast.
Dynamical systems modeling and control are also key ingredients in continuous production and chemical process engineering where we have competence useful for optimizing process performance.
